Hello! My name is Nicholas Ho and I am a third year Computational Biology PhD student in Carnegie Mellon University School of Computer Science. I am incredibly grateful to be advised by Professor Jian Ma and Professor Eric Xing. I develop self-supervised and representation learning models that learn from large biological datasets to uncover new insights across different scales of biology.
Pretraining contextualized representations across biological scales
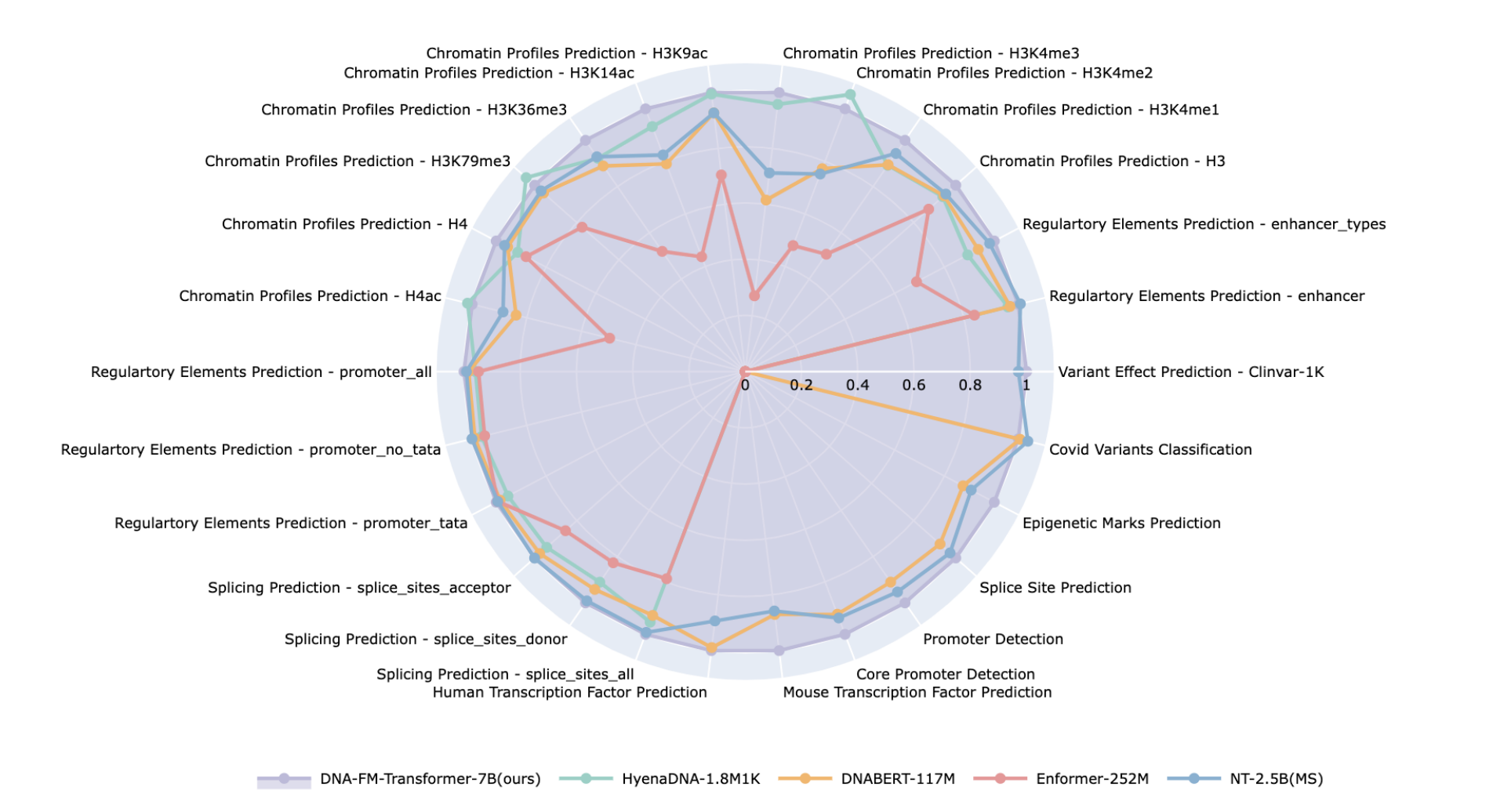
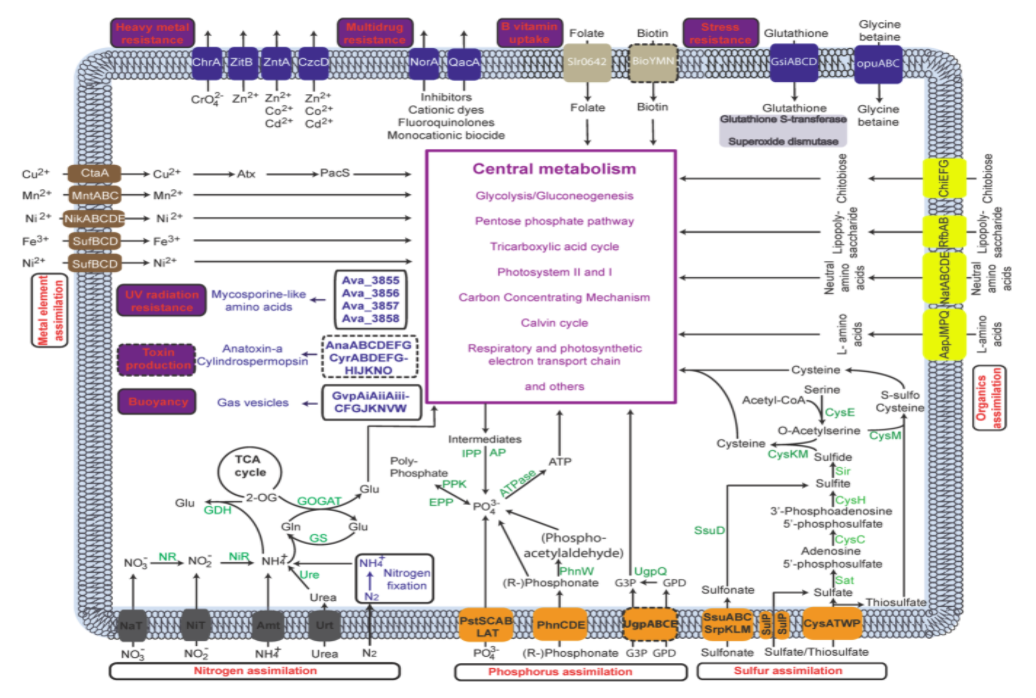
- DNA — Pretrained AIDO.DNA, a 7B parameter genome foundation model trained with 3D parallelism (Megatron-LM), to learn evolutionary representations useful for cis-regulatory sequence-to-function prediction.
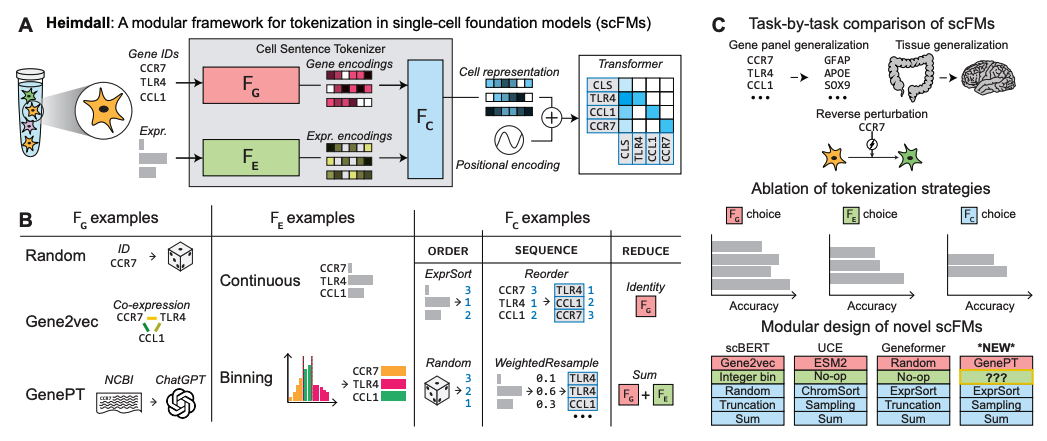
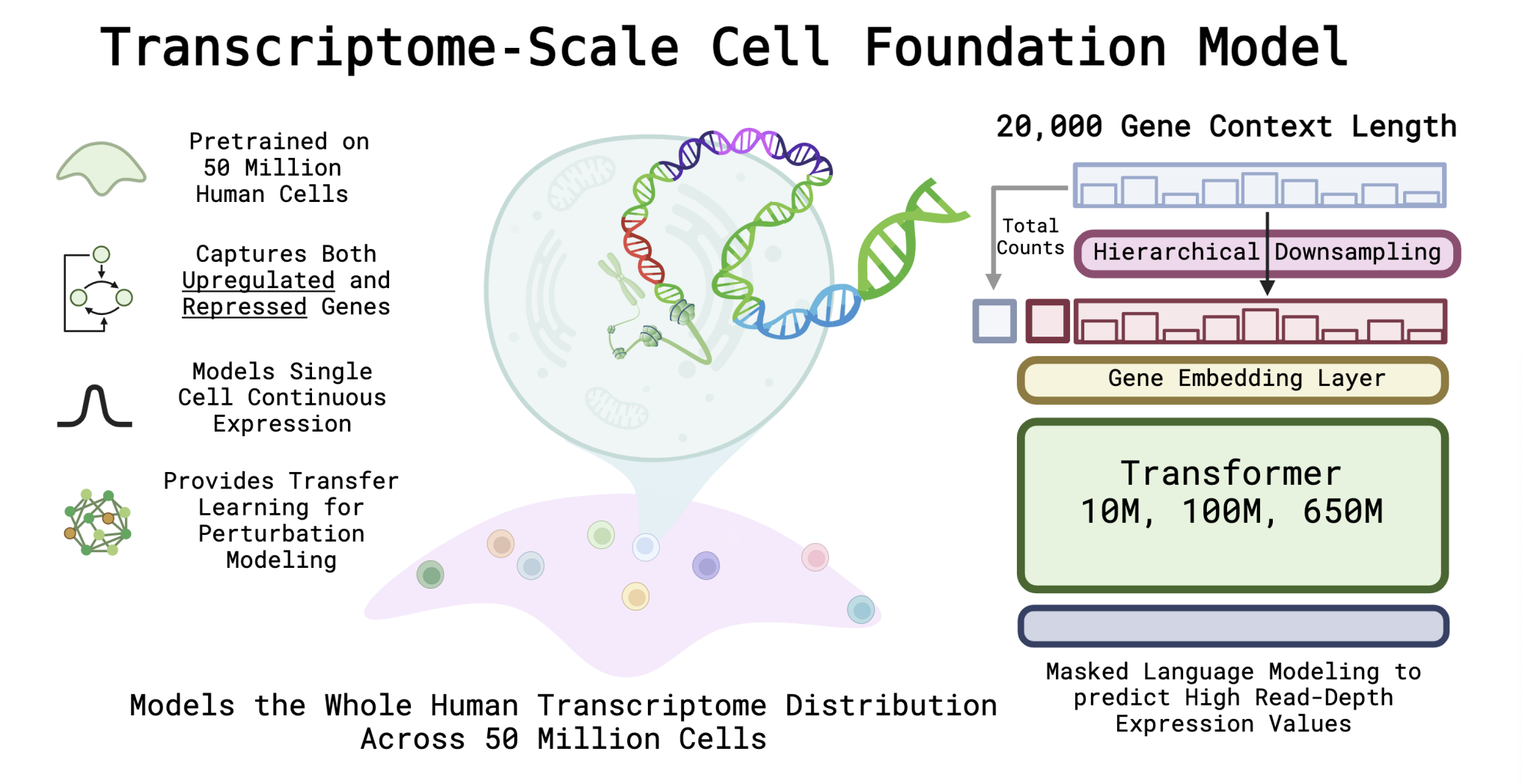
- Single Cell — Built AIDO.Cell, a 600M parameter whole-transcriptome model using masked self-supervised learning, and studied how tokenizer design affects transfer in single-cell foundation models.
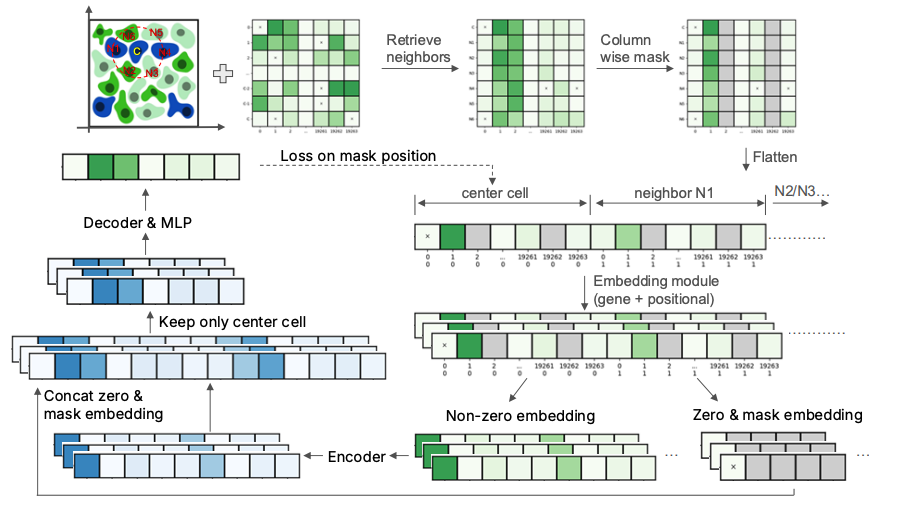
- Tissue — Developed AIDO.Tissue, a spatial transcriptomics foundation model that learns tissue-level context through spatial cell-guided pretraining.
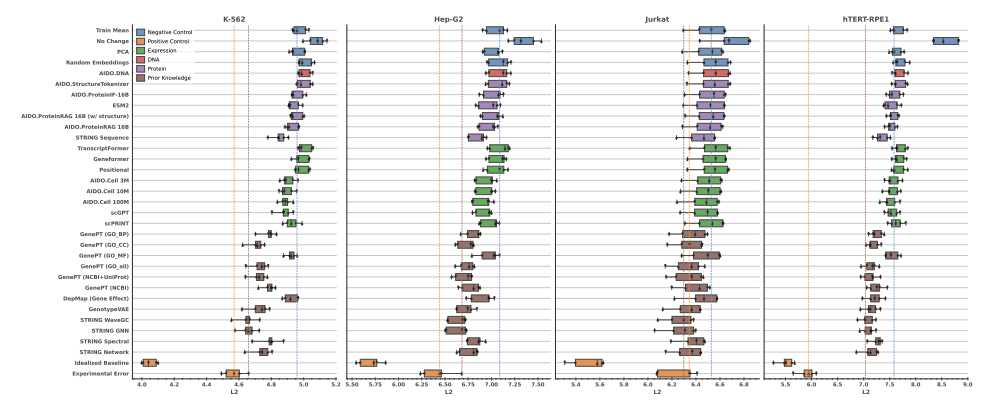
- Perturbation Response — Performed a comprehensive evaluation of perturbation response prediction, training and benchmarking diffusion, flow matching, Schrödinger bridge, and GNN-based models for unseen chemical and genetic perturbations.
I believe that the task always informs the representation. In biology, because data is limited, it is critical to design pretraining objectives aligned with the downstream tasks that matter. By working on the right problems with the right inductive biases, I believe we can build generalizable and scalable models.
Feel free to reach out to me! I'm always looking for interesting problems to talk about!
Publications
Most recent publications on Google Scholar.
‡ indicates equal contribution.
HEIMDALL: Disentangling Tokenizer Design for Robust Transfer in Single-Cell Foundation Models
Ellie Haber*, Shahul Alam*, Nicholas Ho*, Renming Liu, Evan Trop, Shaoheng Liang, Muyu Yang, Spencer Krieger, Jian Ma.
bioRxiv, 2025.
AIDO.Tissue: Spatial Cell-Guided Pretraining for Scalable Spatial Transcriptomics Foundation Model
Jing Gong*, Yixuan Wang*, Nicholas Ho, Xingyi Cheng, Le Song, Eric Xing.
bioRxiv, 2025.
Foundation Models Improve Perturbation Response Prediction
Elijah Cole, Geert-Jan Huizing, Sohan Addagudi, Nicholas Ho, Euxhen Hasanaj, Merel Kuijs, Toby Johnstone, Maria Carilli, Alec Davi, Caleb Ellington, Christoph Feinauer, Pan Li, Romain Menegaux, Shahin Mohammadi, Yanjun Shao, Josiah Zhang, Emma Lundberg, Le Song, Ziv Bar-Joseph, Eric P. Xing.
bioRxiv, 2026.
Scaling Dense Representations for Single Cell with Transcriptome-Scale Context
Nicholas Ho, Caleb N. Ellington, Jinyu Hou, Sohan Addagudi, Shentong Mo, Tianhua Tao, Dian Li, Yonghao Zhuang, Hongyi Wang, Xingyi Cheng, Le Song, Eric P. Xing.
In NeurIPS Workshop on AI for New Drug Modalities, 2024.
Accurate and General DNA Representations Emerge from Genome Foundation Models at Scale
Caleb N. Ellington, Ning Sun, Nicholas Ho, Tianhua Tao, Sazan Mahbub, Dian Li, Yonghao Zhuang, Hongyi Wang, Le Song, Eric P. Xing.
In NeurIPS Workshop on AI for New Drug Modalities, 2024.
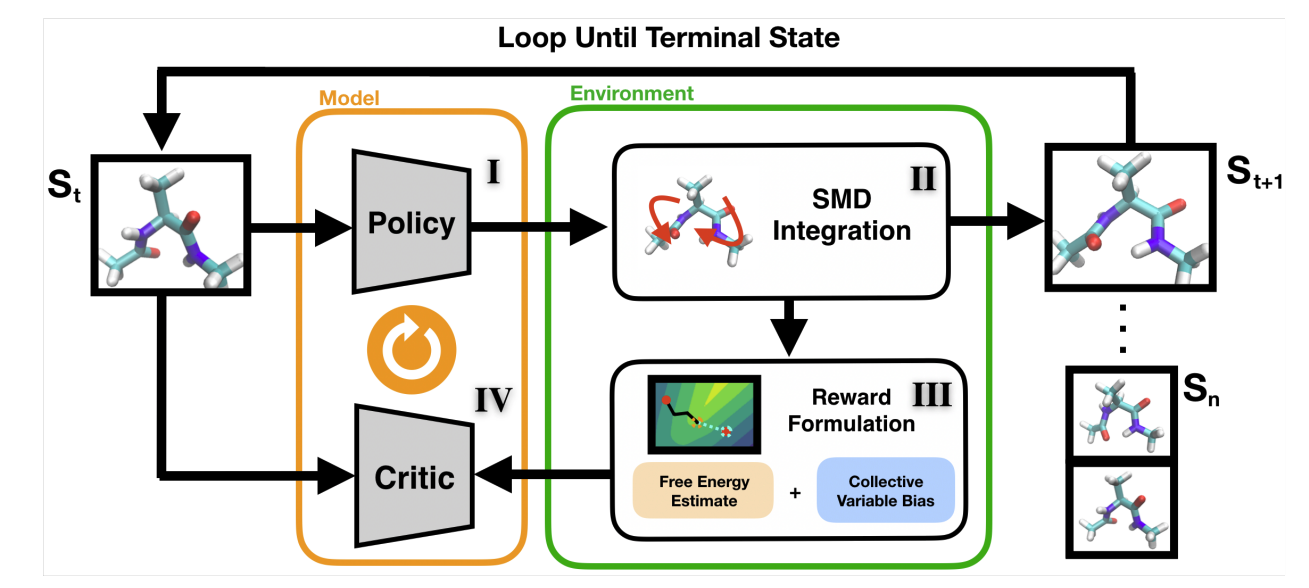
Learning Free Energy Pathways through Reinforcement Learning of Adaptive Steered Molecular Dynamics
Nicholas Ho, John Kevin Cava, John Vant, Ankita Shukla, Jacob Miratsky, Pavan Turaga, Ross Maciejewski, Abhishek Singharoy.
Machine Learning In Structural Biology (MLSB) Workshop at the 36th Conference on Neural Information Processing Systems
HEIMDALL: Disentangling Tokenizer Design for Robust Transfer in Single-Cell Foundation Models
Ellie Haber*, Shahul Alam*, Nicholas Ho*, Renming Liu, Evan Trop, Shaoheng Liang, Muyu Yang, Spencer Krieger, Jian Ma.
bioRxiv, 2025.
AIDO.Tissue: Spatial Cell-Guided Pretraining for Scalable Spatial Transcriptomics Foundation Model
Jing Gong*, Yixuan Wang*, Nicholas Ho, Xingyi Cheng, Le Song, Eric Xing.
bioRxiv, 2025.
Foundation Models Improve Perturbation Response Prediction
Elijah Cole, Geert-Jan Huizing, Sohan Addagudi, Nicholas Ho, Euxhen Hasanaj, Merel Kuijs, Toby Johnstone, Maria Carilli, Alec Davi, Caleb Ellington, Christoph Feinauer, Pan Li, Romain Menegaux, Shahin Mohammadi, Yanjun Shao, Josiah Zhang, Emma Lundberg, Le Song, Ziv Bar-Joseph, Eric P. Xing.
bioRxiv, 2026.
Scaling Dense Representations for Single Cell with Transcriptome-Scale Context
Nicholas Ho, Caleb N. Ellington, Jinyu Hou, Sohan Addagudi, Shentong Mo, Tianhua Tao, Dian Li, Yonghao Zhuang, Hongyi Wang, Xingyi Cheng, Le Song, Eric P. Xing.
In NeurIPS Workshop on AI for New Drug Modalities, 2024.
Accurate and General DNA Representations Emerge from Genome Foundation Models at Scale
Caleb N. Ellington, Ning Sun, Nicholas Ho, Tianhua Tao, Sazan Mahbub, Dian Li, Yonghao Zhuang, Hongyi Wang, Le Song, Eric P. Xing.
In NeurIPS Workshop on AI for New Drug Modalities, 2024.
Learning Free Energy Pathways through Reinforcement Learning of Adaptive Steered Molecular Dynamics
Nicholas Ho, John Kevin Cava, John Vant, Ankita Shukla, Jacob Miratsky, Pavan Turaga, Ross Maciejewski, Abhishek Singharoy.
Machine Learning In Structural Biology (MLSB) Workshop at the 36th Conference on Neural Information Processing Systems
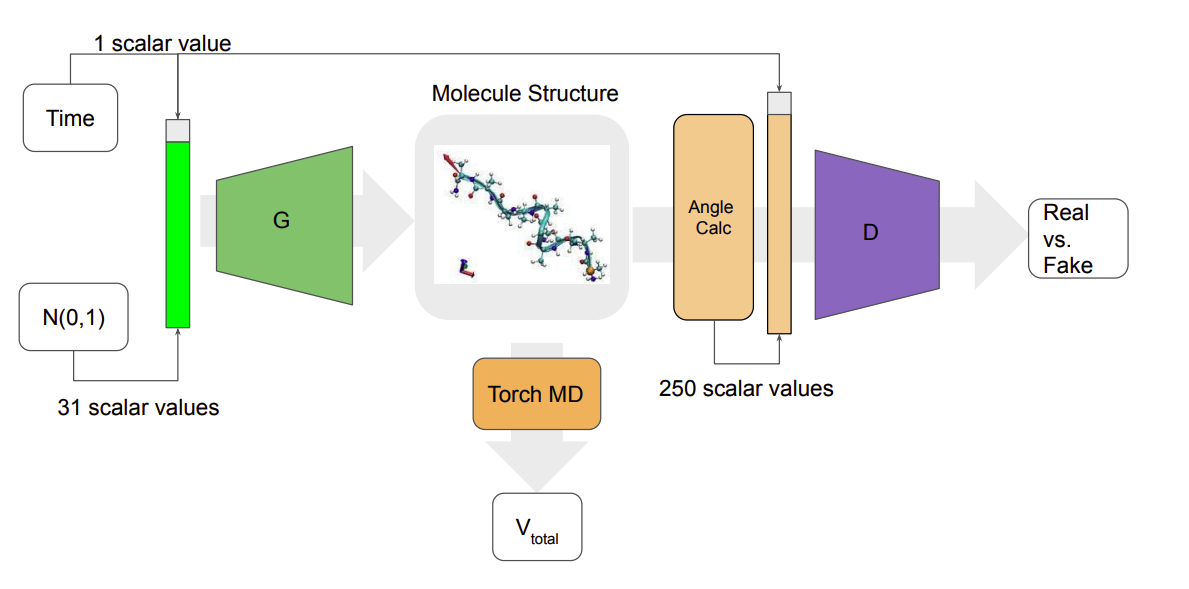
Towards Conditional Generation of Minimal Action Potential Pathways for Molecular Dynamics
John Kevin Cava, John Vant, Nicholas Ho, Ankita Shukla, Pavan Turaga, Ross Maciejewski, Abhishek Singharoy
ELLIS ML4Molecules Workshop, 2021
CyanoPATH: a knowledgebase of genome-scale functional repertoire for toxic cyanobacterial blooms
Wei Du, Gaoyang Li, Nicholas Ho, Landon Jenkins, Drew Hockaday, Jiankang Tan, Huansheng Cao
Briefings in Bioinformatics
Projects
Vitæ
-
Carnegie Mellon University August 2023 - NowPhD Student
Advised by Prof. Jian Ma and Prof. Eric Xing. -
Harvard Medical School June 2022 - May 2023Visiting Research Scholar
• Accepted as a visiting research fellow at Harvard DBMI through a highly competitive application process.
• Currently developing a novel machine learning architecture to deal with missing and imbalanced modalities. -
Struct. Sys. Bio at Biodesign Institute at ASU Feb 2020 - May 2023Research Associate
Worked as a Research Associate in the Singharoy Lab. Lead research for my honors thesis to utilize deep ML methods with statistical mechanical methodologies.
• Lead and wrote a project on discovering free energy pathways with reinforcement learning
• Developed a molecular differentiable simulator for Jax based on TorchMD
• Helped develop novel conditional generative models for deriving free energy pathways
• Implemented and tested several physics-informed models.
-
Carnegie Mellon University June 2021 - August 2021Visiting Research Scholar
Accepted as a visiting research fellow in CMU Statistics through a highly competitive summer research program.
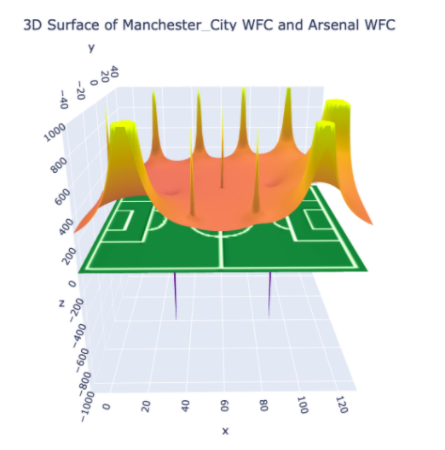
• Created and a novel methodology for using stochastic simulators for soccer game predictions. Our method had comparable results to models trained directly on scores. -
Pichel Lab Biodesign Inst. May 2020 - May 2021Assistant Researcher under Ferran Garcia Pichel at Biodesign Institute
Developed a Python Plugin for Qiime2 for the relationship between ribosomal gene copy number and size. -
Cao Lab Biodesign Inst. August 2019 – May 2021Assistant Researcher
• Conducted data engineering and analysis on genomes assembled from a metagenome in order to study the strain level variance within microbiome communities.
• Implemented a web system that highlighted which genes present in a pathway for particular cyanobacteria species using PFAM’s Hidden Markov Model package, JavaScript, SQL databases, shell and Python. -
Western Tool & Supply June 2020 - July 2020Software Engineer, Intern
• Implemented and trained an LSTM Recurrent Neural Network to predict customer purchase likelihoods. • Integrating and using Bluetooth LE between microcontrollers and Google Chrome into their IOT system. -
TGEN May 2020 - August 2020Virtual Helios Scholar
Due to covid, the Helios Scholars Program was moved to a virtual format. -
Arizona State University August 2019 - May 2023B.Sc. Honors Student
Double Major in Mathematics (4.0) and Computer Science (4.0)
Hobbies
I like to Yoyo!2025 Yoyo Performance (Check this one out)

Thank you Martin Saveski for creating this really neat template!